Genomics Services for Drug Development
Genomics is the science that studies the genome of living beings. Genome studies, performed with technologies such as DNA sequencing via Sanger method or Next Generation Sequencing (NGS), are used to map and study genes and their functions, including the identification of genes responsible for diseases.
New technologies have allowed genomics to explode; Next Generation Sequencing (NGS), genome editing techniques such as CRISPR-Cas9, quantitative technologies to measure gene expression, using qPCR and digital PCR (dPCR).
Genomics services for R&D focus on the study of the genetic code, its regulation, the effects of sequence modifications, and the organisms present in a sample.
This is why Labtoo developed a tailor-made service to support Labs, Biotechs, Medtech, and Pharma companies in their genomics service outsourcing.
Find the right service for your project: NGS and Sanger sequencing, metagenomic and microbiota analyses, genotyping and gene expression by microarray, qPCR, digital PCR (dPCR), molecular biology and cloning, bioinformatics service.
Select your DNA Sequencing
Service
The sequencing of a genome determines the DNA sequence of an organism, either based on a pool of cells or a single cell. Options include Sanger's method for a short sequence, or NGS (Next Generation Sequencing) for more complex sequencing.

Select your RNA Sequencing Service
The transcriptome of an organism refers to the set of messenger RNAs derived from the transcription of the genome. RNA-Seq (RNA sequencing) uses high-throughput sequencing to identify and quantify the mRNA from the genome transcript at a given point in time.

Select your Metagenomics Analysis Service
Metagenomics refers to the genomic study of samples taken from a complex biological environment. This highlights the diversity of microbial ecosystems, whether they are found in marine environments, under the ground, or even in our intestines.

Select your Epigenetics Service
Epigenetics is the study of the mechanisms by which genetic information is reversibly altered. These are induced by the cellular environment, whose signals lead to the regulation of genes. This is of particular interest in the study of the development of diseases that can be caused by epigenetic abnormalities.

Select your Gene Expression Analysis and Microarray Service
Genetic information is not expressed in the same way throughout the life of an organism. Genes can be made active or inactive as needed: this is the principle of gene expression. This expression can be quantified using various techniques that often require a step of amplification of a DNA fragment by PCR (Polymerization Chain Reaction).
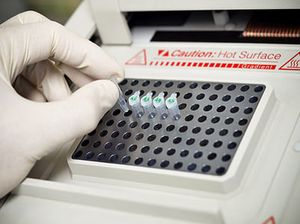
Select your Transgenic Genotyping Service
The genotype is the entire allelic composition of an organism and is responsible for its physical characteristics (phenotype). Genotyping is the analysis of the genetic information of an individual by comparing a DNA sequence with a reference sequence.
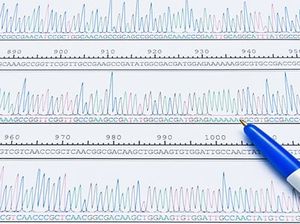
Select your Cloning Service
Molecular cloning allows new genes to be added to the genome of an organism for expression. A DNA sequence of interest and a plasmid are digested and these two sequences are linked. The sequence can be expressed by the plasmid vector. The host organism must be made competent to integrate the plasmid into its genome.

Select your Genome Editing Service
Genome editing corresponds to a genetic modification applicable to any type of cell (animal, plant, microbial etc.). This allows the development of cell models, used for the functional cells. The CRISPR-Cas9 method has revolutionized genome editing with its simplicity and affordability.

Select your Nucleic Acid Sample Preparation Service
The study of the genome requires the preparation of genetic material samples. To this end, protocols for nucleic acid extraction and purification and storage of DNA or RNA samples are useful for genomics projects.

Our team will handle your R&D service management from the beginning to the end
Perform a feasibility study by looking for existing expertise available within the network of partners
Set up a study protocol, financial quotation and preparing contracts with lab partners
Implement the study plan into a schedule, collect all needed materials and reagents and execute the service
Do you need more information about sequencing and genomics services?
How does Next Generation Sequencing (NGS) work?
Today, research instruments and reagents allow us to sequence entire genomes, whether known or not, to focus on regions of interest or to sequence the exome of isolated cells.
How has the sequencing technique developed in the 1970s by the British biochemist Frederick Sanger evolved to enable these advances?
The answer lies in the massively parallel technology offered by Next Generation Sequencing (NGS) and the phenomenal advances in bioinformatics.
Second-generation DNA sequencing is based on parallel amplification. DNA molecules are amplified in an emulsion PCR (emPCR). Adapter ligation and PCR produce DNA libraries with the corresponding 5′ and 3′ ends, which are immobilized on individual oligonucleotide-labeled microbeads. The bead-DNA conjugates can then be emulsified into emulsion droplets containing a single bead.
Clonal amplification then occurs during emPCR, with each template DNA being physically separated from all others, with daughter molecules remaining bound to the microbeads. Alternatively, PCR amplification can produce bridges and form clusters of clonal DNA populations in a solid phase reaction. The libraries produced by these two methods can then be read in a highly parallelized manner: the microbeads produced by emPCR can be washed on a picotiter plate, containing wells large enough to hold a single bead. Flow cell products from bridge amplification can be visualized by detecting fluorescent reversible terminator nucleotides at the ends of an ongoing extension reaction.
Third-generation sequencing is based on single molecule sequencing (SMS), real-time sequencing, or any other feature that varies from second-generation genome sequencing. Two examples of mecanisms:
- Nucleotide detection is done in a Zero Mode Waveguide (ZMW), as in the PacBio sequencers. DNA polymerase molecules are attached to the bottom of each ZMW, and target DNA and fluorescent nucleotides are added. Since the diameter is narrower than the wavelength of the excitation light, the illumination decreases rapidly up the ZMW: only the nucleotides incorporated during polymerization provide bursts of real-time fluorescent signal.
- DNA sequencing of nanopores as used in the MinION sequencer from Oxford Nanopore Technology. Double-stranded DNA is denatured by an enzyme that passes one of the strands (ssDNA) through a biological nanopore embedded in a synthetic membrane, through which a voltage is applied. As the ssDNA passes through the nanopore, the different bases prevent ion flow in a distinct manner, making it possible to deduce the sequence of the molecule by monitoring the current at each channel.
CRISPR-Cas9 and Genome Editing
Since the first publication of the CRISPR mechanism in 1987 by a team from Osaka University, the gene editing method known as CRISPR-Case 9 has come a long way.
What is CRISPR-Cas, how does it work, and what are the steps to do genome editing with this tool?
CRISPR corresponds to a very particular type of DNA sequence of one of the Escherichia coli genes: it consists of 5 short repeated sequences, separated by short unrepeated sequences ("spacers"), hence its name: "clustered regularly inter-spaced short palindromic repeats", or CRISPR.
The Cas genes code for a family of DNA-cutting enzymes. It was described in 2008 that bacteria integrate DNA from viruses at the site of these spacers. As a result of this integration, bacteria acquire the ability to recognize and guide the Cas enzyme to cut viral DNA, and thus deactivate it. In 2012, a team will publish in Science how the CRISPR-Cas9 system can be used to cut any DNA.
The steps for using the method are as follows:
- Select the gene to be modified and obtain its sequence,
- Select the endonuclease to be used: Cas9, Cas12, Cpf1, etc.,
- Construct the RNA Guide (gRNA) using tools such as Benchling,
- Assemble the guide vector in silico, choosing the most appropriate vector, e.g. a vector that has already been used to edit a gene,
- Cloning the guide vector containing: a backbone plasmid, the Cas9 gene, the sequence for the gRNA,
- Insert the vector into the cells to be modified, by transfection, transformation, or viral infection.
The future of technology clearly lays towards this process’ improvement. The technology is still young, both technically (many cases have shown the limits in terms of specificity) and ethically (manipulations on human chimeric embryos are subject to licensing). Genetic engineering actors are developing the next versions of the CRISPR-Cas system.
The different types of providers
Two major categories of genomics service providers:
Academic platforms
Academic platforms are solicited for genome sequencing that requires the development or adaptation of a method. As sequencers are very expensive, academic laboratories may form a group to finance them. The service is provided by a technological platform that made itself available from external structures such as research institutes and companies.
Service companies
Genomics service companies offer services tailored to the demands of private or public clients: DNA sequencing, genome editing, microarray, method development. They can be highly specialized (on a technology or a type of application), or generalist, and can validate methods such as qPCR, using GLP (good laboratory practice) principles.
The post-genomic era
The genomic era is a period that stretches from the discovery of DNA by Watson and Crick to the sequencing of the entire human genome. This project, which began in the late 1980s and ended in 2003, aimed to provide answers to all the questions related to the functioning of our cells.
Far from having met these expectations, the project brought to light a higher level of complexity regarding the mechanisms that regulate the behavior of our cells and the expression of our genes. The era of post-genomic then appeared, bringing with it all the "omics" science groups: proteomics, metabolomics, transcriptomics etc.
Science in the post-genomic era is therefore concerned with determining when and under what conditions a gene is expressed. These studies aim to understand the organization of gene networks, the DNA/DNA, protein/DNA or protein/RNA interactions that lead to the expression of these genes. This era has also allowed the emergence of bioinformatics, whose aim is to model in silico the interaction networks to lead to a virtual living cell.
Used technologies
Molecular cloning techniques
Nucleic acid extraction
PCR, qPCR, dPCR
Sanger method and NGS sequencer
Microarray
CRISPR-Cas9 and other gene-editing technologies

